Reference Genome minutes (Archived)
Minutes for the reference genome meeting, September 26-27, 2007
Orthology determination
Kara Dolinski
Background information
Ortholog tools and model organism database usage
others include:
- HCOP (PMID 16284797),
- treefam (PMID 16381935),
- Compara (no pub yet),
- multiparanoid (multispecies inparanoid) (PMID 16873526)
There are pros and cons with all of the above methods individually. In addition, it is very difficult to assess which is best for the Ref Genome application because none contain the same set of inputs, many use different identifiers, and there is not a clear gold standard that can be used for benchmarking (though there have been some attempts to compare these methods, see PMID 17440619). Another issue is that none contain all of the necessary Ref Genome species. In all cases, it is sometimes necessary to manually look for an ortholog when no tool finds them (short proteins, for example, or divergent, like E, coli proteins. Another problem is the way the model organism databases (MODs) handle orthologs – the mouse schema can’t cope with many to many relationships.
Most of the Ref Genome MODs use YOGY (PMID 16845020), which has all species except chicken and zebra fish – methods include KOGs, InParanoid, homologene, orthoMCL and a table of curated orthologs between budding yeast and fission yeast. An issue with YOGY is that it is not clear that updates and continued maintenance will be happening.
Possible approaches for the Ref Genome Project
- YOGY
One option is to use YOGY because most MODs are already using it. This is dependent upon future maintenance of the project, so the decision was tabled.
- Data integration
One idea is to try a data integration approach, such as one used by Troyanskaya, Roth, Marcotte, Gerstein, etc. For example, the bioPIXIE (PMID 16420673; Princeton), system integrates functional genomics data, with the end result being a probability that two genes are functionally related. We can explore using such an approach with comparative genomics data, where all the different ortholog detection schemes are used as input, the data are integrated, and the result is a probability score.
Pros: 1) We do not need to choose a single method, 2) We do not have to wait for other databases to update the data. Cons: 1) New method for everyone, 2) Need to see how this approach works for comparative genomics data.
We agreed that this option was worth exploring and will be tested by the Princeton groups (Troyanskaya-Dolinski-Botstein).
Action items
- A common set of input protein sets
Suzi and Karen E will generate a page where all sequences will be available.
- All vs. all BLAST
To start, the Princeton group (Kara Dolinski) will run an all vs. all BLAST with the sequences collected above, and make all the data available because at Princeton, the infrastructure, scripts, etc already exist because of the P-POD project (see http://www.ortholog.princeton.edu). We will invite all the ortholog identification groups to take the all v. all BLAST results and run their algorithms. Having them run their algorithms on the same BLAST results should help in assessing the methods.
- Explore data integration approaches
(Kara Dolinski, Olga Troyanskaya, David Botstein as described above)
- Explore using P-POD as a common database of orthologs
The Princeton group offered to put the results of whatever method is chosen into their P-POD database so that curators can search and visualize the orthologous groups
Conclusion
At the end of the day, we need a complete set of orthologs that covers all reference genomes. The orthology determination should capture one-to-many relationships and many-to-many relationships (question: does that need to be captured somewhere? “Unique putative ortholog”,” one2many”, “many2many” (gene family?)). Our concern is to capture the full set rather than making statements about the evolutionary relationships between gene products and/or organisms (we probably need to clarify this in our documentation and what we present to the public as our goals).
Setting curation priorities
Rex Chisholm
When we started the reference genome project last year we made our main priority genes involved in human diseases. The Scientific Advisory Board suggested to also try to curate genes for which there is no GO annotations but that have published data. Other suggestions:
- Encode
- members of a complex should all be done at the same time
- all enzymes in a pathways should be done at the same time
[ACTION ITEM] (everybody): We will add categories of genes to annotate in addition to ‘disease genes’. We will choose five genes from each of the following four groups: 1. diseases (Resources: RGD portal (http://rgd.mcw.edu/dportal/), OMIM (http://www.ncbi.nlm.nih.gov/Omim/getmorbid.cgi) 2. biochemical/ signaling pathways/ (reactome) (Resources: Reactome (http://reactome.org/), Pathway Tools http://bioinformatics.ai.sri.com/ptools/) 3. bleeding edge list: For example: New “hot” genes in your area of interest; genes that come up in computational studies/population studies; most cited papers by some text-mining method; genes cited in newsmedia; newly named genes (HGNC) 4. conserved genes/unannotated genes; genes that have few annotations and have lots of literature. DONE, see documentation here Procedure_for_selection_of_target_genes
[ACTION ITEM] (Val): Provide the list of 207 genes conserved between pombe and human with no annotation/information
[ACTION ITEM] (Jim): Provide the set of conserved genes found by InParanoid that are conserved in all 12 species (660 or so); we might want to prioritize this list by ascending order of number of annotations to target unannotated genes (who can do that?) DONE, see 'Suggestions' spreadsheet (look for the "conserved Hs-Ec" sheet)
[ACTION ITEM] (Ruth): send the HGNC list of genes with few annotations
This will be done on a rotation basis from all databases. I suggest we go alphabetically:
- November 2007: Arabidopsis thaliana
- December 2007: Caenorhabditis elegans
- January 2008: Danio rerio
- February 2008: Dictyostelium discoideum
- March 2008: Drosophila melanogaster
- April 2008: Escherichia coli
- May 2008: Gallus gallus
- June 2008: Homo sapiens
- July 2008: Mus musculus
- August 2008: Rattus norvegicus
- September 2008: Saccharomyces cerevisiae
- October 2008: Schizosaccharomyces pombe
[ACTION ITEM]: contact/meet with people who have made tools for orthology determination to see if they can help us (that possibly includes re-running the analyses using the most recent set of sequences and proper IDs) THIS ACTION ITEM NEEDS TO BE ASSIGNED TO SOMEONE:
- Compara: Emily?
- Homologene: Judy?
- TreeFam
- in paranoid
- others?
[ACTION ITEM]: Kara: run the P-POD over the full ref genomes set? analysis on the ref genome data set. Need computational pipeline with existing resources. Currently takes 3 weeks to do 8 species all v all. Goal was set for February 2008 to include all ref genome sets.
METRICS
The Reference Genome has as one of its goals to provide quantitative measures for annotation progress. Specifically, we want to know the breadth (what fraction of genes have been annotated in each genome) and the depth (to what degree of precision is each gene annotated). This is difficult to do because each measure that we need depends on information that is difficult to measure: (1) total number of genes (and ideally gene products, which includes splice variants), in each organism; (2) number of pseudogenes (which should be excluded when measuring annotation progress); (3) to what granularity a gene may be annotated based on experimental information available. The following presentations addressed different aspects of those issues.
Sequence metrics
Karen Eilbeck Grant has a goal to manage the contents of sequence annotation. Try to address quantitatively, how to evaluate sequence annotations, how different is a genome release from a previous release, complexity of alternative splicing; can we keep track of sequence curation progress? Can we track more than the splits and merges?
- Using GFF3 files, measures 1. sequence annotation turnover, 2. annotation edit distance, 3. splicing complexity
- to specifically measure what was attributable to curation, removed all assembly-induced changed. Only look at annotation where there was no change to underlying.
- measured, between two releases, how many genes still existed in the next release, and how many were traceable to the next release: Worm and c. elegans stable in terms of changes.
- How well does a prediction match a reference annotation? Burset and Guigo 1996 (PMID: 8786136)
- How to compare sequence annotations: based on distance measure (on a scale of 0-1); built on the ideas of sensitivity, specificity and accuracy (referred to as congruency):
- how well does a prediction matches a reference annotation: you can measure true positives, false positive, false negatives (in the bits of the gene models): a bad prediction would have a score of 0, a perfect match would have a score of 1. They give numerical values to sensitivity, specificity and congruency (accuracy). This is also complicated by alternative splicing; they take this into account by comparing each pair of transcripts. See slides for examples or updated gene models and how that affects the scores. From that you get a bar graph that gives a quantitative value of how much the genome has changed between two releases; some genomes like mouse and human vary quite a bit between releases; others (for eg fly) are much more stable.
- Alternative splicing: the trend for alternative splicing is increasing but lower than what is found in the literature. They have a formula to calculate splicing complexity: if the two splice variants are very different from each other they get a higher score (range 0-1). See slides for a graph of the distribution of splicing complexity. One issue is with bad gene models: some may be annotated as splice variants but they could actually be several genes.
- Conservation of alternative splicing: is there any correlation between different species? The answer is that there is a small correlation but not as much as you expect from leading the literature.
- Kimberly points out that Wormbase annotates a mix of genes and proteins. Biggest problem is that protein is often unspecified – don’t know which one it is? Concern about overstating evidence.
How this affect the reference genome project
- Can we use this to implement a mechanism to warn curators that an annotated protein has been modified?
-> Not a high priority since most groups do not annotate to gene products; rather they annotate to genes. It is still the goal to eventually annotate to gene products and we should anticipate potential problems that might create.
- For genes with information in the ‘WITH/FROM’ column, should databases be notified if there is a modification to the sequence corresponding to the ID in the ‘WITH/FROM’ column?
-> The consensus is that although this is important to consider, it’s NOT a priority. When 90% of the genes are annotated we can set that up. Moreover, with the Ref Genomes annotation guidelines, since the WITH is well characterized, this is relatively unlikely to be a major problem (corresponding proteins are expected to be well curated).
- Issues about annotating splice variants
1. For most genes, we are probably not even aware which splice variants exist or are expressed; even when they are known, many papers do not specify which splice variant they use, and many assays do not allow making the distinction. UniProt has a “generic” version of each protein for which there is a splice variant (ie, the complete gene product? or something else more theoretical?), so one can annotate to the generic form when the information is not available. Suzi emphasizes that it’s important to be able to document this and do it to the finest level we can.
2. We are unsure how that affects users that view annotations; using the UniProt IDs it is possible to merge all information to the ‘generic gene’, but we are unsure of whether or not this is true/easy/obvious depending on whether you look at the UniProt page, the QuickGO page, the gene association file. For example, can this ever be computed as annotations to two different genes?
[ACTION ITEM]: (developers/software group): consider the potential impact of annotating to different forms of the gene.
Depth assessement
Chris Mungall Presented different ways to assess depth of annotations. The following graph is an average of each individual measurement (described below) for every database.
ALL annotated genes to the reference genome genes and we can see that the latter have improvements in all categories.

- Distance to leaf
Measures the average maximal distance of annotations relative to a leaf: you take the maximum path of all the terms a gene is annotated to (annotations to the leaf itself get ignored; annotations to root terms get ignored).
Comments:
- Re-calculate with is_a only paths [ACTION ITEM] (Chris)
- Re-calculate with experimental codes only; generate several versions of the data classified by different evidence codes? [ACTION ITEM] (Chris)
- Should annotation to the root or no annotation be considered equivalent? we try to measure *progress*; Chris: there is a technical problem, if there is NO annotations, as far as the database is concerned, that gene does not exist (discussion about the fact that we might want to have the whole set of sequences, etc. No way to deal with this but we should think of ways of addressing this. Loading the total number of genes per organisms would be sufficient?
- Can we use this measure to figure out whether we need to improve the graph? that may point to some genes where annotations need improvement
- Information content
The information content of a node (aka term) in the graph is a measure of specificity derived from the number of annotations to that node. The graph is taken into account, i.e. annotations propagate up the graph to the root. The more annotations there are to a node, the lower the information content. Intuitively, the information content corresponds to the degree of ‘unusualness’ of an annotation to a node. Nodes high in the DAG such as ‘cellular process’ have a low degree of information content because so many annotations are to this term (or one of its children). Nodes deep in the DAG such as ‘recombination nodule’ have high information content because it is unusual for gene products to be annotated to that term. The information content can be a more reliable measure of specificity than depth in the DAG because the DAG is unbalanced (some terms near the root may actually be highly specific). However, it can be prone to annotation bias (for example, immunity-related terms may be unfairly penalized if there is a disproportionately high number of annotations focused in this area). The highest possible value based on the current ontology structure is [15-20?]. The diagram shows the average information content for all gene product annotations.
- DAG Coverage
This measures how many nodes have been annotated to [and sum that up?] (takes into account transitivity). This measures specificity and the breath of annotation for a given gene, ie you read more papers and find a previously un-annotated term in a different area of the graph, this will make the DAG coverage wider.
- Publications per gene
Number of distinct publications associated with a gene through annotations. Suggestion: Re-calculate with only experimental evidence codes since as we curate reference genome gene, we often spend a lot of time replacing NAS and TAS (would be interesting to be able to compare both) [ACTION ITEM] (Chris)
- Term coverage
Number of distinct terms annotated to per gene.
[ACTION ITEM]? (Chris): Provide such reports on a regular basis?
Breadth assessment
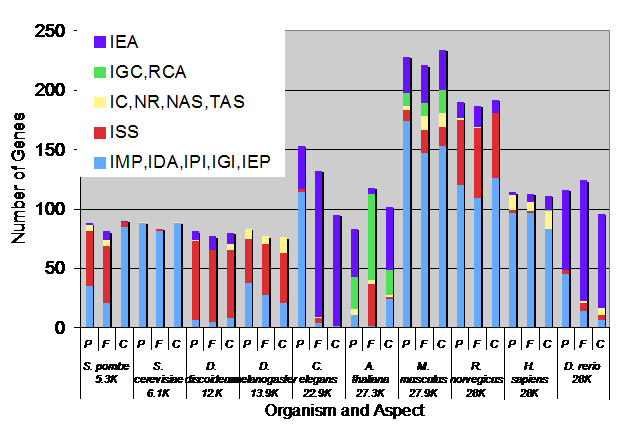
Mike Cherry Data and Figure for this provided by Chris, Susi and Mike provide effective presentation of Breadth assessment. Root annotations are not counted.
- Reference genome genes, Sept 07
- Whole genome, Sept 07
- This indicates that the number of experimental annotations went up and number of IEAs went down
[ACTION ITEM] (Pascale Gaudet): Add to RefGenome curation practices SOPs: please enter your unique gene_id in the google spreadsheet (Makes it easier to parse)
[ACTION ITEM]: (Pascale Gaudet) Generally, provide guidelines for filling the google spreadsheet (IDs, where to put notes, etc)
Counting papers, assessing completeness/comprehensive annotation status
Ruth Lovering Ideally, the completeness of annotation of a gene happens when all papers referring to that gene have been read by a curator (whether or not those papers provided GO new annotations). In practice, however, curating all literature for a gene in only feasible when there are relatively few papers (maximum 5-10). Thus, although we were compiling the number of papers published for a gene, the number of papers read, and the number of papers used for annotation (columns LMNO of the Google spreadsheets), we eventually realized that this number was not always representative of the depth of curation (for example, 20 papers curated out of 1000 is only 5% of the literature, but the gene might be curated to great detail; while 1 paper out of 2 for another gene gives 50% and is probably under-curated). We all agreed that the measures Chris presented are both more meaningful and easier to get (since they can be generated automatically), and therefore we will stop keeping track of the papers.
[ACTION ITEM](Pascale Gaudet): Document in SOPs
Another factor we have been tracking is when a curator judges that the curation of a gene is ‘comprehensive’, that is, that is accurately represents the biology (irrespective of the number of papers available or read). This appears in the spreadsheets. The guideline is that when there are few papers, all papers should be read; when there are many (a curator can judge what is too many), then a review should be read to find the important primary literature and decide what information needs to be captured. We don’t keep track of whether or not reviews have been read. Wormbase uses textpresso (PMID 15383839), that helps ensuring curators do not overlook information. The ‘comprehensive’ curation status doesn’t get invalidated when a newer paper is published; however, curators may (and are encouraged to) update the date when the newer literature is curated.
[ACTION ITEM](Pascale Gaudet): Document in SOPs
Methods
How to balance curation of experimental literature and ISS inference annotations work?
- Some databases (MGI): do not want to concentrate efforts on ISS they prefer to focus on experimental evidence, but other groups (including the SAB) think we are doing the community a disservice by having ND for genes that are well characterized in other organisms. Of course, we don’t all have to have the same annotations. We should try to find out whether we are overlooking too many annotations. Another point was brought up about coexisting annotations (RE: another way to look for omissions) ??
[ACTION ITEM] (Chris): pull out ND annotations and report to each group DONE, see Reference_Genome_Database_Reports
- ISS is done electronically in mice to rat and human based on homologene sets. Co-existence of ND to root AND IEA/ISS? Don’t display ND if ISS is there but ND remains in database. Having IEA and ND in files ok if now splitting files. People thought those should rather be ISS.
Tools
Curation interface demo
Pascale Gaudet Link to tool prototype: http://rails-dev.bioinformatics.northwestern.edu:24000/index.html couple of suggestions, but we reorganized the requirements for the data we collect (mostly papers) so most of the interface will be very different. Some specific suggestions:
- Add Ortholog: Ruth: need search function NOT pull-down for gene symbol. Susan: you enter name and treefam suggestion pops up and suggests orthologs. Eurie: we need mechanisms to download and upload tab-delimited files. * Also: needs to have constraints and checks for existence of primary ID.
- Edit Curation: Pascale: All edits should be on one page.
- Also, organism not just human anymore, now it does not have to be a human ortholog. There are requirements on the wiki, and people should look and add info.
[ACTION ITEM] (Software group): Continue working on development the curation tool
Display in AmiGO
Chris Mungall: Reference genome genes should be highlighted somehow; some people suggest having ‘comprehensive’, ‘in progress’
Graphical display of the annotations
Mary Dolan: The graphs can be viewed at http://www.geneontology.org/images/RefGenomeGraphs/ Main modification made recently is the addition of GO slims in the annotation list (ie, now there are two tables: by the slim and to the exact terms). David suggested using GO term finder to generate a ‘custom’ list of slim terms for those tables? Since the generic GO slim is not great (too general: only contains 150 terms). Mary suggests to show the graphs using slims since the images are very big (creates storage and viewing problems??); many people didn’t like the idea since viewing to slims usually does provide enough information. Chris: we tried this (what did you try? I think collapsing the nodes somehow) for MSH2; not sure about the statistical significance of the data but possibly good. Want check boxes on AmiGo to generate graphs on the fly with and without ISS and with and without slim. Need flags of ref genome to appear in AmiGO. There is also the ability to view the graphs as svg files (on IE?) It was suggested to add colors to the table as displayed in the graph for each organism. Mary suggested to have display for all GO terms like she has for GO slims for each term. She did already provide this, including colors: [http://www.geneontology.org/images/RefGenomeGraphs/5422.html#Slim
[ACTION ITEM] (Mary Dolan, Software group): Continue working on the best way to display annotations graphically.
Annotation consistency
Pascale Gaudet
By annotation consistency I would like to not only address the consistency between curators but also the consistency between the annotations of orthologs across databases. Since those genes are orthologs, we expect some level of conservation of function/process and we can now use the fact that we are annotating the same genes concurrently to provide tools to help make annotations more consistent. Why are orthologous genes annotated differently?
- Different function/process/component in different organisms
- Different experiments done in different systems
- Omission of annotations
- Varying granularity of annotations
- Incorrect annotations
- Problems in the ontology
The first two are acceptable and expected; we would however like to address the other cases.
Omission of annotations
When some genes are annotated to the root, it would be interesting to know whether some information could be obtained via ISS (or were overlooked in the literature).
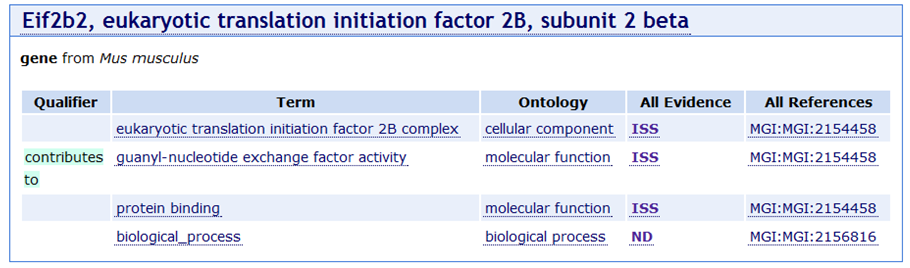
EXAMPLE: It seems intuitively wrong that an initiation factor subunit would be annotated to ‘biological process’. There are concerned that the research community will not understand why such an annotation is missing. David believes this annotation was made by Harold and that he couldn’t find anything to ISS with for the process. There was some discussion about how to deal with this: create links between F and P, use IC more widely. No consensus but the discussion will continue (see later action item about use of ISS, IEA and IC, TAS).
[ACTION ITEM] (Chris) There will be reports of the genes marked ‘comprehensively annotated’ and annotated to the root terms. This will be used as (1) a check to confirm that it was not possible to make a better annotation and also to decide whether or not this really is a problem (maybe it’s always correctly annotated to the root).
DONE, see Reference_Genome_Database_Reports
[ACTION ITEM] (Tanya Berardini, Emily Dimmer, Pascale Gaudet, David Hill, Chris Mungall, Kimberly Van Auken): Write up recommendations for usage of ISS, IEA, IC
Varying granularity of annotations
This can happen when a curator feel he/she doesn’t have the expertise to make an annotation (reading a paper outside their field) and thus chooses to make a more general annotation. It is also possible that a newer term was created since an annotation was made. We do have ways to discuss those (wiki, email list, Source Forge tracker) but those get little usage.
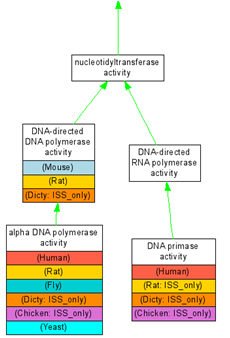
This is may not be the best example. But the basic question is, when such a discrepancy is observed (human gene is annotated to DNA primase, but not human, rat, fly, and yeast), is it real or was some data not captured on some organisms
Incorrect annotations
When one annotation lies in a very far area of the graph compared to all other annotations, a possible reason for this is that the choice of term was incorrect. There are terms for which errors are rather widespread; for example there are many oxidoreductase-type enzymes annotated to ‘electron transport’ because there is an electron transfer during the reaction. Examples: nos1, nos2 (rat), nox1 (human) –nitric oxide synthase-; ADH7 human alcohol dehydrogenase; mouse ReytSat (retinol saturase).
Wrong annotations: Pascale shows example of wrong function annotation in yeast "outlier?": is there a way to use the graph to detect outliers? Mechanism to review outliers: from GO slim list with graphs find outlier on graph - a good measure for quality control. Mary Dolan talked about the confusion matrix they have made to analyze the variations in annotations; maybe we can use this method to identify incorrect annotations?? David points out that misuse of terms indicates problems with definition.
There is an issue with ISS annotations in that some ISS annotations might be left behind when an incorrect one is removed.
Some other varying annotation practices were discussed: when to annotate to binding and when to the complex?
[ACTION ITEM] (all curators): make a list of terms that often have incorrect annotations, so that we can run reports (Chris Mungall?) to double check those annotations. Also in the Annotation Issues Source Forge Tracker, add a category ‘misused terms’ [ACTION ITEM] (SOP group): recommend using the wiki (GOnuts) to add examples of correct use of terms.
Problems in the ontology
Sometimes when annotating a new genome or a new paper, problems in the ontology become apparent. That’s how many changes in the ontology have come about; the question is, is there a way we can use the annotations from the reference genomes to identify such problems. Whatever mechanism we use to deal with #4 (varying granularity of annotations) could lead to suggestions for modifications the ontology.
Outreach
Susan Tweedie Publicity so far:
- talks
- newsletter
- poster at biocurator meeting (Oct 2007)
Needed:
- [ACTION ITEM] Amelia (web page), Susan, Rex, Petra (content), work on web presence:
(The exact contents of this list need to be discussed and defined) - quick link from GO site / involves some (re)writing - summary tables of genes done/in progress - Mary’s graphs with links to AmiGO - Judy: put tabs across the GO page? - Regular gene feature newsletter/web on the front page, or every time you download the page a new gene comes up out of a pool of 10 or so
- Should we give opportunity for feedback: suggestion for changes, comments on annotations
[ACTION ITEM] (who wants this one?) add a link sending to the comments form, add link for comment to featured genes "was this helpful?" comments/suggestions
- AmiGO (longer term)
[ACTION ITEM] (Judy Blake) Contact NCBI/NLM/OMIM to link to reference genome genes
[ACTION ITEM] (Tanya Berardini) Take stack of GO newsletters to Biocurator meeting in October
- Publication:
[ACTION ITEM] (Pascale Gaudet) - WHERE? Genome research, genomics, genome biology -Topics: talk about goals, process, curation priority, ortholog finding, interesting biology (outliers reflect mistakes in annotations or interesting differences in the biology), benefits (improve annotation consistency and ontology quality)
- Genetic Analysis: Model Organisms to Human Biology meeting (San Diego Jan 2008), Is anyone going?
http://www.gsa-modelorganisms.org/
[ACTION ITEM] (Mike Cherry, who else): Organize a reference genome annotation camp, possibly in spring or summer of 2008.