AmiGO Manual: Gene Product Details
Overview
Gene Product Details page provides information about the individual gene product search result. This includes gene product information, with a link to the database providing the data and access to the protein sequence, and all of the Gene Ontology annotations (including GO term, reference, and evidence code) for that gene or gene product.
Information
The information section displays the following data associated with the gene or gene product:
Name
The common name of the gene or gene product (eg. nuclease, ATPase).
Type
The feature or sequence type of the gene or gene product. Typically a gene or protein.
Species
The species from which the gene product comes. The species name is hyperlinked to the NCBI Taxonomy Browser, which provides more information on the selected species.
Synonyms
Other names for the gene or gene product.
Database
The data source that provided the annotations to the GO Consortium.
Sequence
Clicking the View Sequence hyperlink will display the sequence of the gene product in FASTA format. The FASTA header contains all available identifiers for that sequence. Clicking on the Use as BLAST query sequence will submit this sequence to the GO BLAST server.
Term Associations
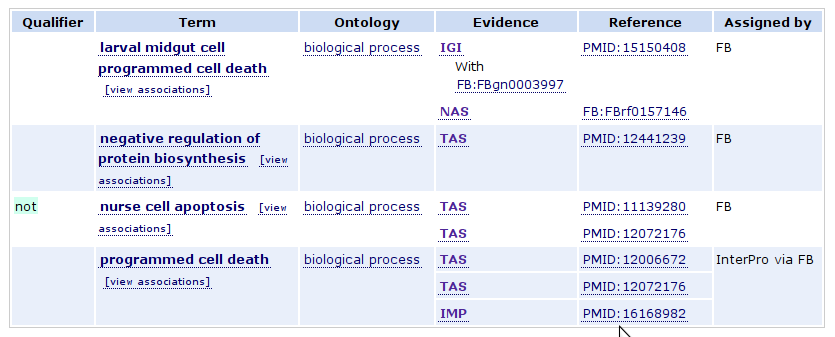
Term associations displays a table of the GO terms that the selected gene or gene product has been associated.
Qualifier
The Qualifier column is used for flags that modify the interpretation of an annotation. Allowable values are NOT, contributes_to, and colocalizes_with. See the GO annotation guide for more information.
Term name
Clicking the term name will display the AmiGO_Manual:_Term_Details; clicking the view association hyperlink will display the AmiGO_Manual:_Term_Associations, all the gene products associated with the selected term.
Ontology
Denotes the ontology that the term belongs to. Clicking the ontology name will display the GO documentation on that ontology.
Evidence code
Every GO annotation (or term association) must indicate the type of evidence that supports it; these evidence codes correspond to broad categories of experimental or other support. Mouse over the evidence code to see the full name of an evidence code. Clicking the selected evidence code will display the evidence code documentation page where detailed description of the selected evidence code can be found.
Reference
Clicking the reference hyperlink will display the reference used to make the association.
Assigned by
The database or group that made the annotation is listed. If this is different to the database from which the gene or gene product comes, it will state that the data were submitted "via" another data source.
Filtering
Term associations can be filtered by ontology and evidence code to limit the terms shown to those in the selected ontology (or ontologies) that are annotated with the chosen evidence code(s). Multiple options can be selected in all the filters; select more than one option by control-clicking (apple-click on a Mac) the names. Click Set filters to set your display options. If no options are selected or you click Remove all filters, AmiGO will display all associations.