Guide to GO Evidence Codes: Difference between revisions
No edit summary |
mNo edit summary |
||
| (32 intermediate revisions by 2 users not shown) | |||
| Line 1: | Line 1: | ||
= Introduction = | |||
These guidelines are a guide to standard usage of the GO evidence codes. | |||
A GO annotation consists of a GO term associated with a specific reference that describes the work or analysis upon which the association between a specific GO term and gene product is based. Each annotation must also include an evidence code to indicate how the annotation to a particular term is supported. Although evidence codes do reflect the type of work or analysis described in the cited reference which supports the GO term to gene product association, they are not necessarily a classification of types of experiments/analyses. Note that these evidence codes are intended for use in conjunction with GO terms, and should not be considered in isolation from the terms. If a reference describes multiple methods that each provide evidence to make a GO annotation to a particular term, then multiple annotations with identical GO identifiers and reference identifiers but different evidence codes may be made. | A GO annotation consists of a GO term associated with a specific reference that describes the work or analysis upon which the association between a specific GO term and gene product is based. Each annotation must also include an evidence code to indicate how the annotation to a particular term is supported. Although evidence codes do reflect the type of work or analysis described in the cited reference which supports the GO term to gene product association, they are not necessarily a classification of types of experiments/analyses. Note that these evidence codes are intended for use in conjunction with GO terms, and should not be considered in isolation from the terms. If a reference describes multiple methods that each provide evidence to make a GO annotation to a particular term, then multiple annotations with identical GO identifiers and reference identifiers but different evidence codes may be made. | ||
Evidence codes are '''not''' statements of the quality of the annotation. Within each evidence code classification, some methods produce annotations of higher confidence or greater specificity than other methods, in addition the way in which a technique has been applied or interpreted in a paper will also affect the quality of the resulting annotation. Thus evidence codes '''cannot''' be used as a measure of the quality of the annotation. | |||
Evidence codes fall into eight general categories as described below. | |||
Annotators may also find the [http://wiki.geneontology.org/index.php/Guide_to_GO_Evidence_Codes#Evidence_codes_Decision_tree evidence code decision tree] useful in selecting the correct evidence code for an annotation. | |||
= Experimental Evidence Codes = | |||
Use of an experimental evidence code in a GO annotation indicates that the cited paper displayed results from a physical characterization of a gene or gene product that has supported the association of a GO term. The '''Experimental Evidence Codes''' are: | Use of an experimental evidence code in a GO annotation indicates that the cited paper displayed results from a physical characterization of a gene or gene product that has supported the association of a GO term. The '''Experimental Evidence Codes''' are: | ||
*[ | *[http://wiki.geneontology.org/index.php/Inferred_from_Experiment_(EXP) Inferred from Experiment (EXP), (ECO:0000269)] | ||
*[ | *[http://wiki.geneontology.org/index.php/Inferred_from_Direct_Assay_(IDA) Inferred from Direct Assay (IDA), (ECO:0000314)] | ||
*[ | *[http://wiki.geneontology.org/index.php/Inferred_from_Physical_Interaction_(IPI) Inferred from Physical Interaction (IPI), (ECO:0000353)] | ||
*[ | *[http://wiki.geneontology.org/index.php/Inferred_from_Mutant_Phenotype_(IMP) Inferred from Mutant Phenotype (IMP), (ECO:0000315)] | ||
*[ | *[http://wiki.geneontology.org/index.php/Inferred_from_Genetic_Interaction_(IGI) Inferred from Genetic Interaction (IGI), (ECO:0000316)] | ||
*[ | *[http://wiki.geneontology.org/index.php/Inferred_from_Expression_Pattern_(IEP) Inferred from Expression Pattern (IEP), (ECO:0000270)] | ||
= High Throughput Experimental Evidence Codes = | |||
High throughput (HTP) evidence codes may be used to make annotations based upon high throughput methodologies. Use of HTP evidence codes should be carefully considered and follow the GOC's guidelines for their use. The '''High Throughput Evidence Codes''' are: | High throughput (HTP) evidence codes may be used to make annotations based upon high throughput methodologies. Use of HTP evidence codes should be carefully considered and follow the GOC's guidelines for their use. The '''High Throughput Experimental Evidence Codes''' are: | ||
*[ | *[http://wiki.geneontology.org/index.php/Inferred_from_High_Throughput_Experiment_(HTP) Inferred from High Throughput Experiment (HTP), (ECO:0006056)] | ||
*[ | *[http://wiki.geneontology.org/index.php/Inferred_from_High_Throughput_Direct_Assay_(HDA) Inferred from High Throughput Direct Assay (HDA), (ECO:0007005)] | ||
*[ | *[http://wiki.geneontology.org/index.php/Inferred_from_High_Throughput_Mutant_Phenotype_(HMP) Inferred from High Throughput Mutant Phenotype (HMP), (ECO:0007001)] | ||
*[ | *[http://wiki.geneontology.org/index.php/Inferred_from_High_Throughput_Genetic_Interaction_(HGI) Inferred from High Throughput Genetic Interaction (HGI), (ECO:0007003)] | ||
*[ | *[http://wiki.geneontology.org/index.php/Inferred_from_High_Throughput_Expression_Pattern_(HEP) Inferred from High Throughput Expression Pattern (HEP), (ECO:0007007)] | ||
Use of the | = Similarity Evidence Codes = | ||
Use of the similarity evidence codes indicates that the annotation is based on an in silico analysis of the gene sequence described in the cited reference. The evidence codes in this category also indicate a varying degree of curatorial input. The '''Similarity Evidence Codes''' are: | |||
*[ | *[http://wiki.geneontology.org/index.php/Inferred_from_Sequence_or_structural_Similarity_(ISS) Inferred from Sequence or structural Similarity (ISS), (ECO:0000250)] | ||
*[ | *[http://wiki.geneontology.org/index.php/Inferred_from_Sequence_Orthology_(ISO) Inferred from Sequence Orthology (ISO), (ECO:0000266)] | ||
*[ | *[http://wiki.geneontology.org/index.php/Inferred_from_Sequence_Alignment_(ISA) Inferred from Sequence Alignment (ISA), (ECO:0000247)] | ||
*[ | *[http://wiki.geneontology.org/index.php/Inferred_from_Sequence_Model_(ISM) Inferred from Sequence Model (ISM), (ECO:0000255)] | ||
*[ | *[http://wiki.geneontology.org/index.php/Inferred_from_Genomic_Context_(IGC) Inferred from Genomic Context (IGC), (ECO_0000317)] | ||
*[ | *[http://wiki.geneontology.org/index.php/Inferred_from_Biological_aspect_of_Ancestor_(IBA) Inferred from Biological aspect of Ancestor (IBA), (ECO:0000318)] | ||
*[ | *[http://wiki.geneontology.org/index.php/Inferred_from_Biological_aspect_of_Descendant_(IBD) Inferred from Biological aspect of Descendant (IBD), (ECO:0000319)] | ||
*[ | *[http://wiki.geneontology.org/index.php/Inferred_from_Key_Residues_(IKR) Inferred from Key Residues (IKR), (ECO:0000320)] | ||
*[ | *[http://wiki.geneontology.org/index.php/Inferred_from_Rapid_Divergence(IRD) Inferred from Rapid Divergence (IRD), (ECO:0000321)] | ||
= Combinatorial Evidence Codes = | |||
*[http://wiki.geneontology.org/index.php/Inferred_from_Reviewed_Computational_Analysis_(RCA) Inferred from Reviewed Computational Analysis (RCA), (ECO:0000245)] | |||
= Author Statements = | |||
Author statement codes indicate that the annotation was made on the basis of a statement made by the author(s) in the reference cited. The '''Author Statement Evidence Codes''' used by GO are: | Author statement codes indicate that the annotation was made on the basis of a statement made by the author(s) in the reference cited. The '''Author Statement Evidence Codes''' used by GO are: | ||
*[ | *[http://wiki.geneontology.org/index.php/Traceable_Author_Statement_(TAS) Traceable Author Statement (TAS), (ECO:0000303)] | ||
*[ | *[http://wiki.geneontology.org/index.php/Non-traceable_Author_Statement_(NAS) Non-traceable Author Statement (NAS), (ECO:0000303)] | ||
= Curator Inference = | |||
Use of the curatorial statement evidence codes indicates an annotation made on the basis of a curatorial judgement that does not fit into one of the other evidence code classifications. The '''Curatorial Statement Evidence Codes''' are: | Use of the curatorial statement evidence codes indicates an annotation made on the basis of a curatorial judgement that does not fit into one of the other evidence code classifications. The '''Curatorial Statement Evidence Codes''' are: | ||
*[ | *[http://wiki.geneontology.org/index.php/Inferred_by_Curator_(IC) Inferred by Curator (IC), (ECO:0000305)] | ||
= No Biological Data = | |||
*[http://wiki.geneontology.org/index.php/No_biological_Data_available_(ND)_evidence_code No biological Data available (ND), (ECO:0000307)] | |||
= Automatic Assertion = | |||
GO also used one evidence code that is assigned by automated methods, without curatorial judgement. The '''Automatically-Assigned Evidence Code''' is: | |||
*[http://wiki.geneontology.org/index.php/Inferred_from_Electronic_Annotation_(IEA) Inferred from Electronic Annotation (IEA), (ECO:0000501)] | |||
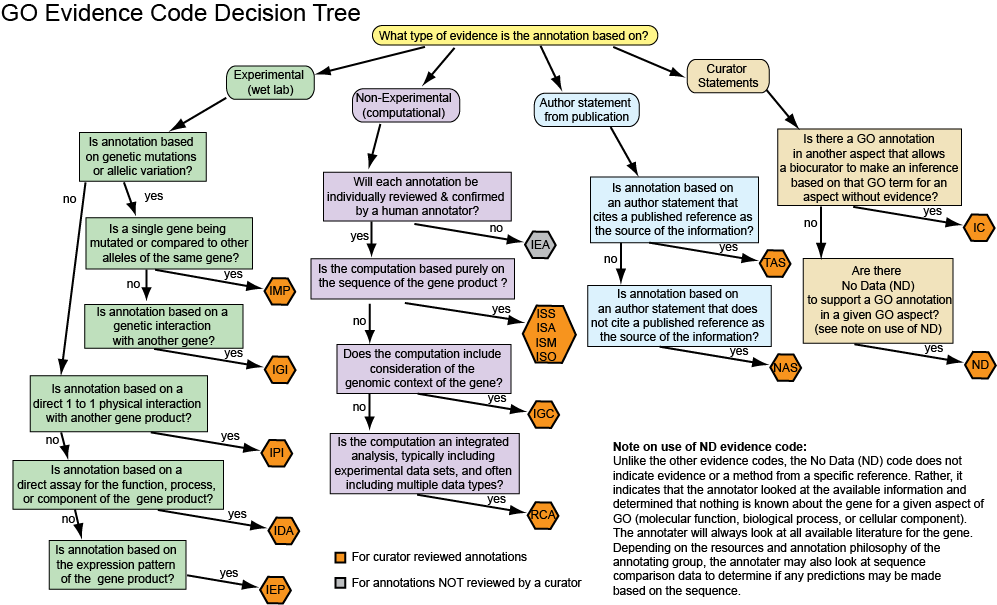
=Evidence codes Decision tree= | |||
[[File:Diag-evCodeFlowChartGood.png]] | |||
== Review Status == | |||
Last reviewed: February 23, 2018 | |||
[[Category:Evidence Codes]] | [[Category:Evidence Codes]] | ||
Revision as of 11:41, 13 April 2019
Introduction
These guidelines are a guide to standard usage of the GO evidence codes.
A GO annotation consists of a GO term associated with a specific reference that describes the work or analysis upon which the association between a specific GO term and gene product is based. Each annotation must also include an evidence code to indicate how the annotation to a particular term is supported. Although evidence codes do reflect the type of work or analysis described in the cited reference which supports the GO term to gene product association, they are not necessarily a classification of types of experiments/analyses. Note that these evidence codes are intended for use in conjunction with GO terms, and should not be considered in isolation from the terms. If a reference describes multiple methods that each provide evidence to make a GO annotation to a particular term, then multiple annotations with identical GO identifiers and reference identifiers but different evidence codes may be made.
Evidence codes are not statements of the quality of the annotation. Within each evidence code classification, some methods produce annotations of higher confidence or greater specificity than other methods, in addition the way in which a technique has been applied or interpreted in a paper will also affect the quality of the resulting annotation. Thus evidence codes cannot be used as a measure of the quality of the annotation.
Evidence codes fall into eight general categories as described below.
Annotators may also find the evidence code decision tree useful in selecting the correct evidence code for an annotation.
Experimental Evidence Codes
Use of an experimental evidence code in a GO annotation indicates that the cited paper displayed results from a physical characterization of a gene or gene product that has supported the association of a GO term. The Experimental Evidence Codes are:
- Inferred from Experiment (EXP), (ECO:0000269)
- Inferred from Direct Assay (IDA), (ECO:0000314)
- Inferred from Physical Interaction (IPI), (ECO:0000353)
- Inferred from Mutant Phenotype (IMP), (ECO:0000315)
- Inferred from Genetic Interaction (IGI), (ECO:0000316)
- Inferred from Expression Pattern (IEP), (ECO:0000270)
High Throughput Experimental Evidence Codes
High throughput (HTP) evidence codes may be used to make annotations based upon high throughput methodologies. Use of HTP evidence codes should be carefully considered and follow the GOC's guidelines for their use. The High Throughput Experimental Evidence Codes are:
- Inferred from High Throughput Experiment (HTP), (ECO:0006056)
- Inferred from High Throughput Direct Assay (HDA), (ECO:0007005)
- Inferred from High Throughput Mutant Phenotype (HMP), (ECO:0007001)
- Inferred from High Throughput Genetic Interaction (HGI), (ECO:0007003)
- Inferred from High Throughput Expression Pattern (HEP), (ECO:0007007)
Similarity Evidence Codes
Use of the similarity evidence codes indicates that the annotation is based on an in silico analysis of the gene sequence described in the cited reference. The evidence codes in this category also indicate a varying degree of curatorial input. The Similarity Evidence Codes are:
- Inferred from Sequence or structural Similarity (ISS), (ECO:0000250)
- Inferred from Sequence Orthology (ISO), (ECO:0000266)
- Inferred from Sequence Alignment (ISA), (ECO:0000247)
- Inferred from Sequence Model (ISM), (ECO:0000255)
- Inferred from Genomic Context (IGC), (ECO_0000317)
- Inferred from Biological aspect of Ancestor (IBA), (ECO:0000318)
- Inferred from Biological aspect of Descendant (IBD), (ECO:0000319)
- Inferred from Key Residues (IKR), (ECO:0000320)
- Inferred from Rapid Divergence (IRD), (ECO:0000321)
Combinatorial Evidence Codes
Author Statements
Author statement codes indicate that the annotation was made on the basis of a statement made by the author(s) in the reference cited. The Author Statement Evidence Codes used by GO are:
Curator Inference
Use of the curatorial statement evidence codes indicates an annotation made on the basis of a curatorial judgement that does not fit into one of the other evidence code classifications. The Curatorial Statement Evidence Codes are:
No Biological Data
Automatic Assertion
GO also used one evidence code that is assigned by automated methods, without curatorial judgement. The Automatically-Assigned Evidence Code is:
Evidence codes Decision tree
Review Status
Last reviewed: February 23, 2018