Noctua
GO-CAMs and Noctua
- This documentation is presented in two parts:
- GO-CAM Principles
- Noctua Curation Tool
GO-CAM Principles
Standard GO Annotations and GO-CAM Models
Standard GO Annotations
GO annotations have been a key component of GO since its inception. Standard annotations are defined as an association between a gene and a biological concept from one of the three GO aspects: Molecular Function (MF), Biological Process (BP), and Cellular Component (CC). Standard annotations always contain a reference (either a published, peer-reviewed paper or internal GO reference) and an evidence code that indicates the type of experiment or method used to make the assertion. Standard GO annotations may further be qualified using annotation extensions that provide additional biological context to a GO term using a relation from the Relations Ontology (RO) and a term from GO or an external ontology, e.g. UBERON.
GO-CAM Models
While standard GO annotations are very useful for discerning basic information about genes, they provide only a partial view of each gene's role in a larger biological context. To provide more comprehensive annotation of genes and link their activities in a causal framework, the GO developed GO-CAMs. Using RO relations, GO-CAMs link GO annotations together with biological entities and external ontology terms to model how a gene functions in the broader context of a biological process or pathway. GO-CAMs thus provide structured descriptions of biological systems and allow for interrogation of causal events in biology through use of clearly defined, and consistently applied, semantics.
A summary of the GO-CAM model specifications is presented in Figure 1.
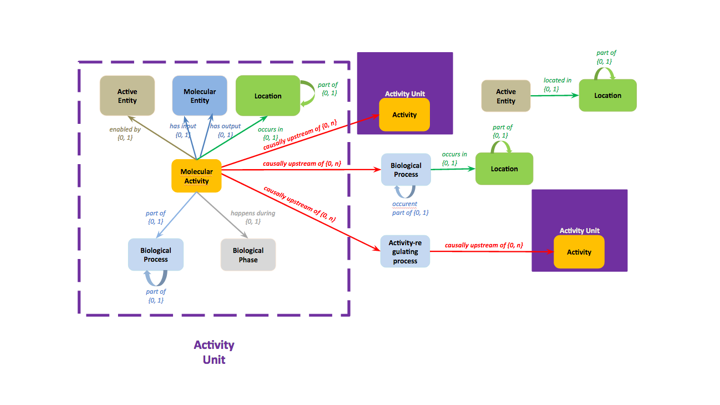
The basic unit of a GO-CAM model is the Activity Unit, outlined on the left, which represents a set of standard GO annotations with select annotation extensions, e.g. the inputs and outputs of a molecular function. GO-CAM models are constructed by filling in as many pieces of relevant information in an Activity Unit as possible and then linking different Activity Units in a causal chain to model a biological process. Thus, GO-CAM models use standard GO annotations as the foundation on which to build more comprehensive representations of biology.
Use of GO in GO-CAMs
Molecular Activities in GO-CAMs
- Wherever possible, curators should strive to select the single most granular GO Molecular Function (MF) term that best describes the overall activity of the gene, gene product, or protein-containing complex being annotated.
- If desired, individual "sub-functions" may be captured by using the 'part of' relation between the main MF and its sub-functions.
Biological Processes in GO-CAMs
- The ultimate aim of GO-CAMs is to create a suite of Biological Process (BP)-centric models that can be used to interrogate causal effects of molecular activities on one another as part of the execution of a larger BP.
- When annotating, curators should always think about the BP they are modeling and what MFs are 'part of' that BP.
- Additional relations between MFs and BPs, e.g. 'causally upstream of or within', can be used to capture experimental information that, in the future, will be incorporated into a more complete model of that process.
Cellular Components in GO-CAMs
- Cellular component annotations are intended to capture where the MF enabled by a gene, gene product, or protein-containing complex occurs.
- Cellular component annotations may be further qualified with cell, tissue, and organismal context if that information is germane to the process being modeled.
Noctua: the Gene Ontology's GO-CAM Annotation Tool
Noctua is a web-based, collaborative Gene Ontology (GO) annotation tool developed by the GO Consortium. Noctua can be used to create standard GO annotations as well as more expressive models of biological processes, known as GO-CAMs (Gene Ontology Causal Activity Models). There are two types of user interface available in Noctua: 1) a form interface and 2) a graph interface.
System Requirements
Noctua is a web-based annotation tool and thus requires only a web browser to access and use.
We have tested Noctua primarily with Chrome on a Mac operating system.
Issues that arise using other browsers and operating systems may be reported on go-helpdesk
User Account Setup
GO-CAMs can be browsed using Noctua, but no annotations can be created or edited unless a user has a registered account.
To create a new account, please email help@geneontology.org or enter a ticket on the helpdesk repo in github.
Note that to create a Noctua account, you will need an ORCID and a github account.
Please allow 24 hours for your account to be created.
If you have any questions about user accounts, please contact help@geneontology.org
Entities and Ontologies for Annotation
Entities for Annotation
Genes and Gene Products
- The primary nodes in a GO-CAM model are the ACTIVITIES (Molecular Functions (MFs)) of genes, gene products, or protein-containing complexes.
- Every gene, gene product, and protein-containing complex annotated in GO-CAMs must be associated with a stable database identifier and represented either in a GPI (Gene Production Information) file (preferred), in an existing annotation file, e.g. a GAF (Gene Association File), or in the GO Cellular Component aspect.
- Curators should strive to annotate activities (MFs) to individual genes or gene products wherever possible.
Protein-Containing Complexes
- In GO-CAMs, curators should always try to assign each member of a protein-containing complex its specific activity (e.g. regulatory activity vs catalytic activity).
- However, if the main activity of the protein-containing complex cannot be ascribed to a single subunit of the complex (e.g. RNA polymerase II activity), then the activity should be enabled by an appropriate protein-containing complex (e.g. a GO protein-containing complex), with each gene or gene product associated with that protein-containing complex with a 'part of' relation.
- Requests to add new entity identifiers to Noctua should be directed to help@geneontology.org
Ontologies for Annotation
Gene Ontology
- The GO is used to annotate:
- Molecular Activities (MF)
- Biological Processes (BP)
- Cellular Component (CC)
- To provide appropriate biological context to an annotation or an activity unit, additional ontologies may be used either in GO-CAM or in annotation extensions [link].
Relations Ontology
Relations used in GO-CAM are a subset of Relations Ontology. These relations and their usage are described in the Annotation Relations page.
Cell and Anatomy Ontologies
- Describes the location where processes and functions occur.
- Describes the location of a GO cellular component.
- Add list
Biological Phase and Life Stage Ontologies
- Describes the temporal period during which processes and functions occur.
- Describes the temporal period during which a cellular component or anatomical entity exists.
- Add list
Chemical Ontology (ChEBI) [link]
- Can be used to capture inputs and outputs of processes and functions.
- GO-CAM uses the Chemical Entities of Biological Interest (ChEBI)
- Sequence Ontology [link]
Requests to add ontologies to Noctua should be sent to help@geneontology.org
GO-CAM Workflow
- The ultimate goal for GO-CAMs is to create a knowledge graph whereby users can use the GO to traverse a causal representation of a biological system.
- To that end, curators should try, as much as possible, to make individual annotations in the context of the overall process being modeled.
- It can be very helpful to refer to a summary figure from a recent research article or review to help visualize a potential GO-CAM.
- When making a GO-CAM model, we suggesting these steps:
- What are the main activities (MFs) of each of the gene products in a model?
- How do those activities relate, in a causal chain, to each other?
- What processes are those activities involved in?
- Where do the activities occur?
- Even when annotating a single paper, try to incorporate as much of this workflow as possible. This will make it easier, in the future, to build on existing models with new curation.
Noctua Users Manual
- Noctua is accessed via: http://noctua.geneontology.org/
Noctua Landing Page
- The Noctua landing page is the portal by which curators can browse or search and filter models.
- It is also the starting point for curation (when logged in) and where individual GPAD and OWL files for a model can be downloaded.
- By default, the Noctua landing page displays models by date, descending order, i.e. the most recently edited models are shown at the top of the list.

Filtering Models
- There are two ways to filter GO-CAMs on the landing page:
- Click on the magnifying glass icon in the upper left
- Click on the metadata icons to the right of the model title in the table.

Filtering with the Magnifying Glass
Clicking on the magnifying glass opens up the menu of available filter options:
- Ontology term (autocomplete)
- Gene product (autocomplete)
- Reference
- If entered as free-text, must be the full reference id, e.g. PMID:31884020 or doi:10.1016/j.ydbio.2019.12.010
- Can also use the drop-down prefix menu (and PMID look-up feature) by clicking on the =+ icon.
- Must press return after entering search string.
- Organism (drop-down list)
- Contributor (autocomplete and drop-down list)
- Groups (autocomplete and drop-down list)
- Exact date (enter YYYY-MM-DD/return or calendar, select/return)
- Date range (enter YYYY-MM-DD/return or calendar, select/return)
- Title (enter/return)
- State (drop-down list)

Noctua Form and Graph Editors
Noctua has three curation interfaces: 1) the Noctua Visual Editor, 2) the Noctua Form and 3) the Noctua Graph Editor.
- The Noctua Visual Editor (VPE) is designed for curating GO-CAM models, i.e. models that include at least two activities/MFs linked by causal relations.
- The Noctua Form is a structured annotation form that is recommended for creating 'standard' GO annotations.
- The Noctua Graph Editor is best suited for linking standard annotations together to create a causal model (GO-CAM).
| Noctua Users Manual | ||
|---|---|---|
| Getting started | Browsing and searching annotations and models | |
| Login | ||
| Creating standard GO annotations | ||
| Form Editor | Molecular Function | |
| Biological Process | ||
| Cellular Component | ||
| Graph Editor | Molecular Function | |
| Biological Process | ||
| Cellular Component | ||
| Adding contextual information (annotation extensions) | ||
| Form Editor | Molecular Function | |
| Biological Process | ||
| Cellular Component | ||
| Graph Editor | Molecular Function | |
| Biological Process | ||
| Cellular Component | ||
| Editing annotations | ||
| Form Editor | ||
| Graph Editor | ||
| Creating GO-CAMs | ||
| Creating an activity unit | Form Editor | |
| Graph Editor | ||
| Linking Activities | Form Editor | |
| Graph Editor | ||
| Model metadata | ||
| Model titles | General Guidelines | |
| Form Editor | ||
| Graph Editor | ||
| Releasing models to production | Form Editor Form Editor] | |
| Graph Editor | ||
| Other tips and tricks | Adding a NOT qualifier to an annotation | |
| Importing existing annotations | ||
| Changing annotation group | ||
| Model validation | ||
| Running the reasoner | ||
| Viewing GPAD export] | ||
| Using templates | ||
Noctua Annotation Review
Documentation on how to use the Noctua Annotation Review workbench is here.
Model Copy
- Noctua has a model copy functionality that allows users to copy an entire model, minus the evidence.
- Model copy can help curators make efficient use of existing models to create new content.
- More details on model copy and how to use it can be found here.
Noctua Maintenance
- The Noctua curation tool undergoes routine maintenance on the second and fourth Thursdays of each month from ~4pm - 7pm PDT.
- During these maintenance outages the following tasks are typically performed:
- Ontology updates:
- incorporate the latest versions of all ontologies used in Noctua, including NEO (Noctua Entity Ontology)
- Model updates:
- replace obsolete ontology terms if there is a 'replaced by' value
- replace ontology terms if usage has changed
- delete any models marked with state 'delete'
- Software updates:
- Incorporate bug fixes
- Add new features
- Ontology updates:
- Noctua maintenance outage reminders are sent out on the go-consortium mailing list ~24 hrs. prior to the outage.
- For each outage, a list of specific tasks is enumerated in a ticket on the noctua github repo.