Ref Gen pub draft (Retired): Difference between revisions
No edit summary |
|||
| Line 27: | Line 27: | ||
==Graphic Representation== | ==Graphic Representation== | ||
[[image:POLAgraph.png]] | A colorful graphic representation of the reference genome effort can be viewed in full here:<br> | ||
http://www.geneontology.org/images/RefGenomeGraphs/ <br> | |||
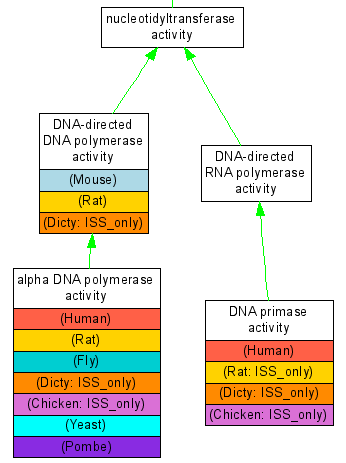
There is one graph per curated reference gene. In addition to the graph there are two informative tables per gene; these list GO annotations by category, and full experimantal annotations in each organism. This facilitates the comparison of the curation status in the 12 reference genomes and helps curators to identifty genes that need attention. | |||
[[image:POLAgraph.png]] Partial Graph of Gene POLA | |||
Revision as of 11:51, 30 October 2007
The Reference Genome Annotation Project
Introduction
The GO consortium has established the complete annotation of 12 reference genomes as a priority goal. These reference genomes are:
- Arabidopsis thaliana (http://www.arabidopsis.org/)
- Caenorhabditis elegans (http://www.wormbase.org/)
- Danio rerio (zebrafish; http://zfin.org)
- Dictyostelium discoideum (http://www.dictybase.org/)
- Drosophila melanogaster (http://flybase.org/)
- Escherichia coli (http://www.tigr.org/)?
- Gallus gallus (http://www.agbase.msstate.edu/)
- Homo sapiens (http://www.ebi.ac.uk/GOA/human_release.html)?
- Mus musculus (http://www.informatics.jax.org/)
- Rattus norvegicus (http://rgd.mcw.edu/)
- Saccharomyces cerevisiae (http://www.yeastgenome.org/)
- Schizosaccharomyces pombe (http://www.genedb.org/genedb/pombe/)
The Reference Genome GO Annotation Team, with representatives from each genome annotation group, will coordinate annotation, facilitate implementation of GO Consortium annotation priorities, provide metrics to assess progress toward the goal of broad and deep annotation of the reference genomes. This group will be responsible for the coordination of the annotation of the twelve reference genomes. This group represents the annotation expertise within the GO consortium and provides key liaisons to the model organism databases the have primary responsibilities for the annotation of the reference genomes.
Purpose
Activities
Graphic Representation
A colorful graphic representation of the reference genome effort can be viewed in full here:
http://www.geneontology.org/images/RefGenomeGraphs/
There is one graph per curated reference gene. In addition to the graph there are two informative tables per gene; these list GO annotations by category, and full experimantal annotations in each organism. This facilitates the comparison of the curation status in the 12 reference genomes and helps curators to identifty genes that need attention.