Ref Gen pub draft (Retired): Difference between revisions
No edit summary |
|||
| Line 39: | Line 39: | ||
*Where they exist, identify the ortholog(s)/homolog(s) of the selected target genes in each species [ at present each database uses their own criteria to decide which orthologous/homologous genes to curate - (include link to these?? on a separate page. Note they are not in a standard format at present but could maybe be summarised - if we decide to include these then each group should review content to make sure it is ok for the public site - I know mine is a bit informal at present) but we are investigating a standardised approach. s.t. <i> I would delete all in the bracket and we put a question for the group here, like 'should we ommit any mentioning of orthology selection until we have a standardised procedure?'</i> pf] | *Where they exist, identify the ortholog(s)/homolog(s) of the selected target genes in each species [ at present each database uses their own criteria to decide which orthologous/homologous genes to curate - (include link to these?? on a separate page. Note they are not in a standard format at present but could maybe be summarised - if we decide to include these then each group should review content to make sure it is ok for the public site - I know mine is a bit informal at present) but we are investigating a standardised approach. s.t. <i> I would delete all in the bracket and we put a question for the group here, like 'should we ommit any mentioning of orthology selection until we have a standardised procedure?'</i> pf] | ||
*Enter the gene identifiers in a shared spreadsheet so that all curators can see the set of genes being curated. | *Enter the gene identifiers in a shared spreadsheet so that all curators can see the set of genes being curated. | ||
*Collect and annotate available literature about the genes | *Collect and annotate available literature about the genes. | ||
*Assign GO terms based on experimental data. | *Assign GO terms based on experimental data. | ||
*Review existing GO annotations to make sure they conform to agreed [[#How does this project differ from standard GO annotation?| standards]] | *Review existing GO annotations to make sure they conform to agreed [[#How does this project differ from standard GO annotation?| standards]]. | ||
*Record in shared spreadsheet that GO annotation when is considered comprehensive for each gene | *Record in shared spreadsheet that GO annotation when is considered [[How do we know when GO annotation is comprehensive?| comprehensive]] for each gene. | ||
A web tool for reference genome annotation is under development. This will facilitate keeping track of and comparing annotations. | |||
==How does this project differ from standard GO annotation?== | ==How does this project differ from standard GO annotation?== | ||
| Line 55: | Line 57: | ||
==How do we know when GO annotation is comprehensive?== | ==How do we know when GO annotation is comprehensive?== | ||
The amount of literature per gene is very variable. Where possible we review every paper about a given gene and capture all possible GO terms but this is only feasible when there are tens of papers. For genes associated with hundreds or even thousands of publications we cannot read all of the papers so we do our best to prioritise the literature and capture all aspects of gene with GO terms. In this situation we often work from recent reviews to lead us to key experimental papers. | The amount of literature per gene is very variable. Where possible we review every paper about a given gene and capture all possible GO terms but this is only feasible when there are tens of papers. For genes associated with hundreds or even thousands of publications we cannot read all of the papers so we do our best to prioritise the literature and capture all aspects of the gene with GO terms. In this situation we often work from recent reviews to lead us to key experimental papers. Users are encouraged to notify us if we have failed to capture some aspect of a specific gene [go help - <i>or do we need a separate forum?</i>]. | ||
In some cases, there is no experimental data for any of the reference genome species but experimental data may be available in other model systems; in these cases we submit GO annotations for the relevant species to GOA [link] so that this information is captured from the primary literature. | In some cases, there is no experimental data for any of the reference genome species but experimental data may be available in other model systems; in these cases we submit GO annotations for the relevant species to GOA [link] so that this information is captured from the primary literature. | ||
| Line 63: | Line 65: | ||
All GO annotations from this project are included in the [[http://www.geneontology.org/GO.current.annotations.shtml| gene association files]] that each group submits to GO and can be viewed using [[http://www.geneontology.org/amigo/help-term_assoc.shtml| AmiGO]]. | All GO annotations from this project are included in the [[http://www.geneontology.org/GO.current.annotations.shtml| gene association files]] that each group submits to GO and can be viewed using [[http://www.geneontology.org/amigo/help-term_assoc.shtml| AmiGO]]. | ||
It is also possible to view the | It is also possible to specifically view the reference genome effort. A list of all annotated genes linking to colorful graphics can be viewed in full here:<br> | ||
http://www.geneontology.org/images/RefGenomeGraphs/ <br> | http://www.geneontology.org/images/RefGenomeGraphs/ <br> | ||
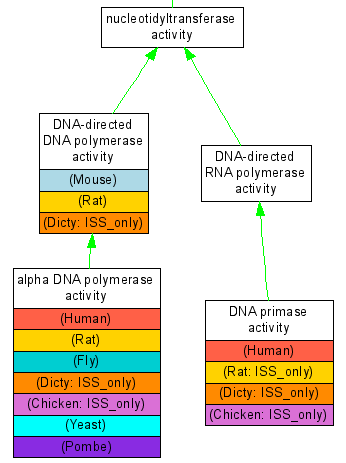
Each curated reference gene links to one graph. In addition to the graph each page includes two informative tables, which either lists GO annotations by broad GO category (GO slim), or shows full experimental annotations in each organism for the given gene. This facilitates comparison of the curation status in the 12 reference genomes and helps curators to identify genes that need attention. | |||
[[image:POLAgraph.png]] <b>Partial Graph of Gene POLA</b> | [[image:POLAgraph.png]] <b>Partial Graph of Gene POLA</b> | ||
==Activities== | ==Activities== | ||
(this is probably not necessary?) | (<i>this is probably not necessary for the public site?</i>) | ||
* Monthly Conference Calls | * Monthly Conference Calls | ||
* First Reference Genome Annotation Meeting, Princeton, NJ, Sept 26, 27, 2007 | * First Reference Genome Annotation Meeting, Princeton, NJ, Sept 26, 27, 2007 | ||
=Questions for ref genome call on 11/13/07= | |||
* Dicuss orthology determination here or wait until we have standardised procedure? | |||
* Provide a section about 'metrics', or is that rather just for internal use? (showing a graph or part of the graph from Chris that's on the [[http://gocwiki.geneontology.org/index.php/Reference_Genome_minutes#Depth_assessement| ref genome minutes page]] might be attractive? | |||
* We discussed at meeting having a 'comments' form or something. | |||
* Do we need the Activities section? | |||
Revision as of 15:04, 5 November 2007
The Reference Genome Annotation Project
Introduction
We are experiencing an explosion of genomic information, with more and more genomes being sequenced and ever growing genomic data. However, there are limited resources to annotate these growing numbers of genomes, thus automatic annotation will be the method of choice for many. On the other hand, all model organism databases have a group of trained and highly skilled GO curators. In an effort to maximize and optimize the GO annotation of a large and representative set of key genomes (called from now on 'the reference genomes') the GO consortium has established the priority goal to completely annotate 12 reference genomes. These reference genomes are:
- Arabidopsis thaliana (http://www.arabidopsis.org/)
- Caenorhabditis elegans (http://www.wormbase.org/)
- Danio rerio (zebrafish; http://zfin.org)
- Dictyostelium discoideum (http://www.dictybase.org/)
- Drosophila melanogaster (http://flybase.org/)
- Escherichia coli (http://www.tigr.org/)?
- Gallus gallus (http://www.agbase.msstate.edu/)
- Homo sapiens (http://www.ebi.ac.uk/GOA/human_release.html)?
- Mus musculus (http://www.informatics.jax.org/)
- Rattus norvegicus (http://rgd.mcw.edu/)
- Saccharomyces cerevisiae (http://www.yeastgenome.org/)
- Schizosaccharomyces pombe (http://www.genedb.org/genedb/pombe/)
The Reference Genome GO Annotation Team, with representatives from each genome annotation group, coordinates annotation, facilitates implementation of GO Consortium annotation priorities, and provides quantitative measures to assess progress toward the goal of broad and deep annotation of the reference genomes. This group represents the annotation expertise within the GO consortium and provides key liaisons to the model organism databases that have primary responsibilities for the annotation of the reference genomes.
Priorities for Annotation
Our ultimate aim is to provide comprehensive GO annotation for all gene products in each of the reference genomes. This is a huge task and requires to prioritise the targets for curation. Our initial annotation efforts (Aug 2006- Sept 2007) focussed on orthologs of human disease genes but in Oct 2007 we widened our list to four priority areas:
- Orthologs of human disease genes
- Topical or ‘hot’ genes
- Genes conserved from E. coli to human that currently lack GO annotation
- Genes involved in (metabolic?) pathways
Each month we curate 5 genes from each category as selected by one of the participating databases on a rotational basis.
Overview of project strategy
Every month each database curates the same set of 20 genes from our priority list. Working on the same genes together promotes discussion about annotations across groups and frequently leads to new terms being added to the gene ontology.
Curation process summary:
- Where they exist, identify the ortholog(s)/homolog(s) of the selected target genes in each species [ at present each database uses their own criteria to decide which orthologous/homologous genes to curate - (include link to these?? on a separate page. Note they are not in a standard format at present but could maybe be summarised - if we decide to include these then each group should review content to make sure it is ok for the public site - I know mine is a bit informal at present) but we are investigating a standardised approach. s.t. I would delete all in the bracket and we put a question for the group here, like 'should we ommit any mentioning of orthology selection until we have a standardised procedure?' pf]
- Enter the gene identifiers in a shared spreadsheet so that all curators can see the set of genes being curated.
- Collect and annotate available literature about the genes.
- Assign GO terms based on experimental data.
- Review existing GO annotations to make sure they conform to agreed standards.
- Record in shared spreadsheet that GO annotation when is considered comprehensive for each gene.
A web tool for reference genome annotation is under development. This will facilitate keeping track of and comparing annotations.
How does this project differ from standard GO annotation?
The reference genome databases have agreed to follow guidelines that are more stringent than those used for standard annotation:
- Experimental evidence codes (IDA, IMP, IGI, IPI, IEP) should be used where possible
- Terms inferred from sequence and structural similarity (ISS) should only be used where the terms are supported by experimental evidence for the similar sequence
- Non-traceable author statements (NAS) should be avoided
- No new annotations should be based on traceable author statements (TAS); existing terms assigned with TAS should gradually be replaced with the appropriate experimental evidence code based on the primary literature
How do we know when GO annotation is comprehensive?
The amount of literature per gene is very variable. Where possible we review every paper about a given gene and capture all possible GO terms but this is only feasible when there are tens of papers. For genes associated with hundreds or even thousands of publications we cannot read all of the papers so we do our best to prioritise the literature and capture all aspects of the gene with GO terms. In this situation we often work from recent reviews to lead us to key experimental papers. Users are encouraged to notify us if we have failed to capture some aspect of a specific gene [go help - or do we need a separate forum?].
In some cases, there is no experimental data for any of the reference genome species but experimental data may be available in other model systems; in these cases we submit GO annotations for the relevant species to GOA [link] so that this information is captured from the primary literature.
Where can GO annotations from the project be viewed?
All GO annotations from this project are included in the [gene association files] that each group submits to GO and can be viewed using [AmiGO].
It is also possible to specifically view the reference genome effort. A list of all annotated genes linking to colorful graphics can be viewed in full here:
http://www.geneontology.org/images/RefGenomeGraphs/
Each curated reference gene links to one graph. In addition to the graph each page includes two informative tables, which either lists GO annotations by broad GO category (GO slim), or shows full experimental annotations in each organism for the given gene. This facilitates comparison of the curation status in the 12 reference genomes and helps curators to identify genes that need attention.
Activities
(this is probably not necessary for the public site?)
- Monthly Conference Calls
- First Reference Genome Annotation Meeting, Princeton, NJ, Sept 26, 27, 2007
Questions for ref genome call on 11/13/07
- Dicuss orthology determination here or wait until we have standardised procedure?
- Provide a section about 'metrics', or is that rather just for internal use? (showing a graph or part of the graph from Chris that's on the [ref genome minutes page] might be attractive?
- We discussed at meeting having a 'comments' form or something.
- Do we need the Activities section?