Obol:Browser
Introduction
Obol-web is an interface to the Obol term decomposition system. The interface is also a prototype platform for new amigo or oboedit presentation paradigms
For more details on Obol, see
The interface can be accessed at
Guided Tour
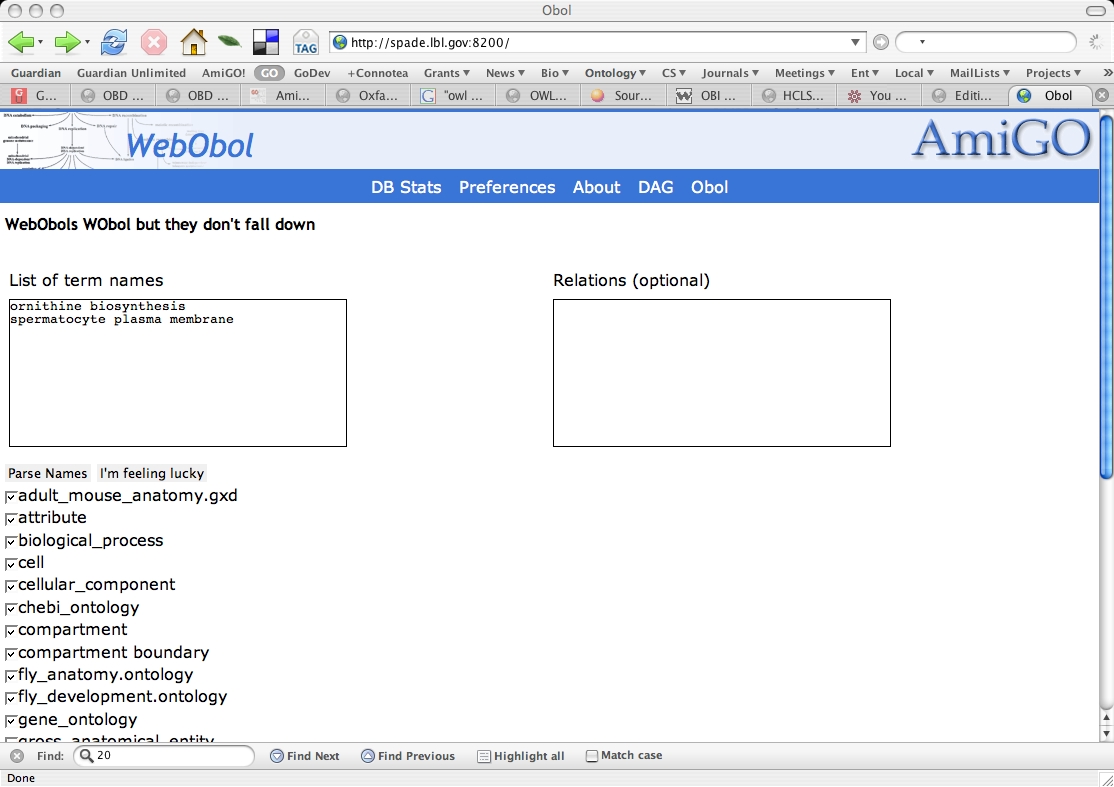
Obol intro screen
The intro screen lets you type in various terms to be decomposed. The screenshot shows two terms ready to be decomposed - ornithine biosynthesis and spermatocyte plasma membrane.
Parse choices
We select "parse" and are taken to a screen showing a selection of parses for each term. The first has an unambiguous parse, the second has multiple parses. The best guess is pre-selected::
Parses selected
We choose the two decompositions, and they are loaded into the (temporary) Obol database. We can see the obo format export (this can be pasted directly into an existing obo file). Note the 'intersection_of' lines. Since "spermatocyte plasma membrane" did not previously exist as a term, a new termporary ID has been granted (note: the Obol database is not persistent, this will be lost when the server restarts!)
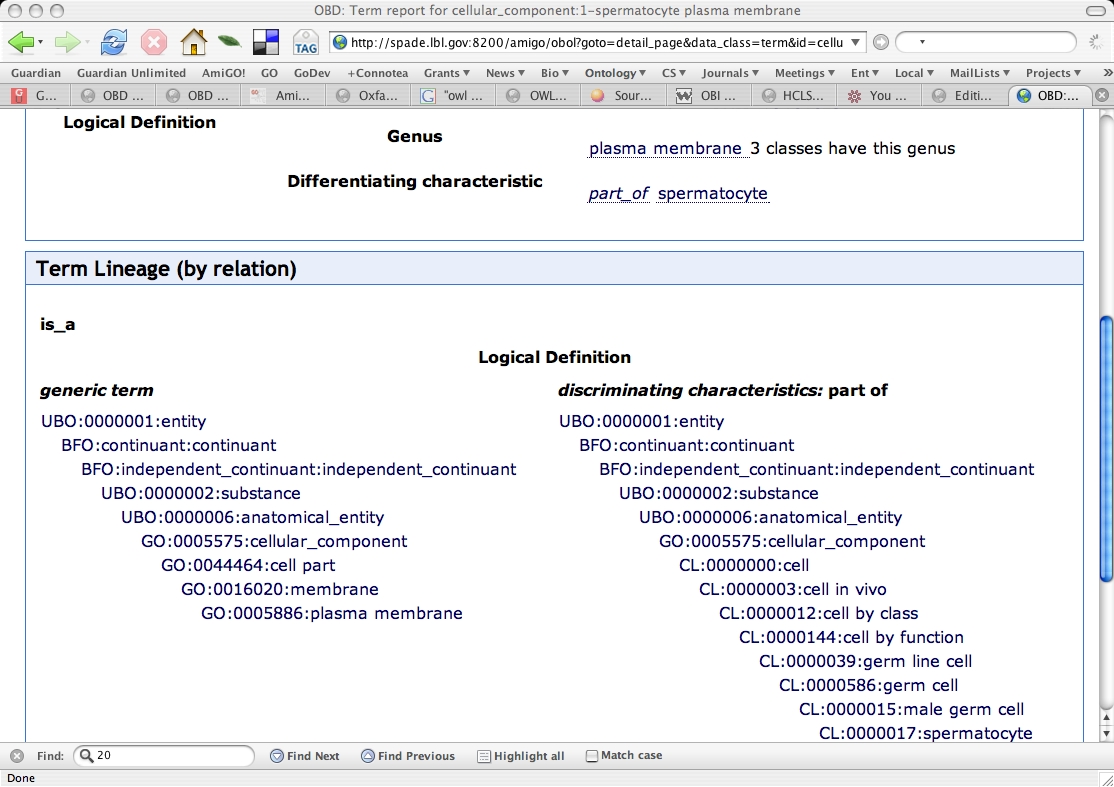
Genus-differentia display in browser
We click on one of the terms to see how it is displayed in the Obol term lineage display. We see distinct lineages for the genus and differentia:
Tree Explorer
Click on DAG to get a tree browser. This has several foundry ontologies loaded. Note that nodes such as "biological process" are placed under the appropriate upper ontology nodes (the upper ontology is required for Obol to do its decomposition and reasoning). You can type a term string into the search box; the screenshot shows "neuron" in the textbox
Search results and filter
Here are the search results for "neuron" filtered to biological_process
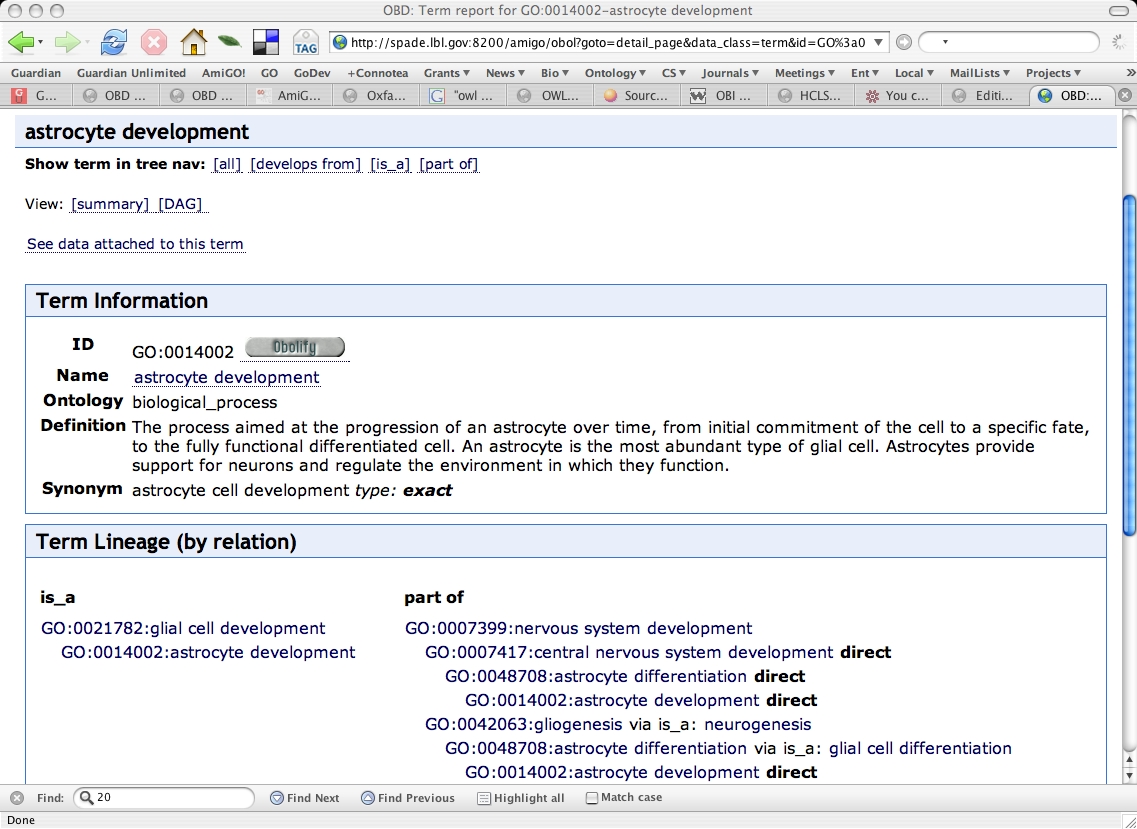
Term detail page
Term details for "astrocyte development". Note the "obolify" button. We can press this to decompose the term
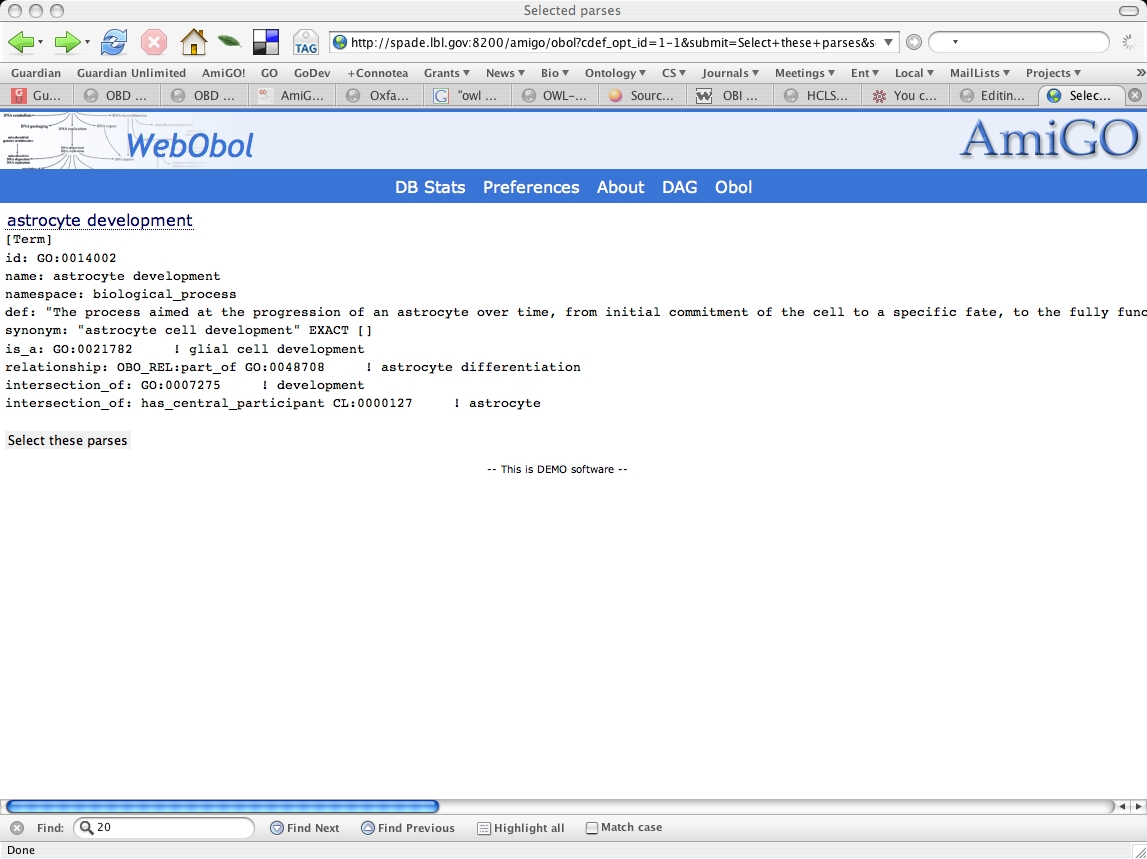
Decomposition for astrocyte development
After selecting the most likely parse for this term, we see the definition in obo format. The logical definition has been loaded into the Obol database
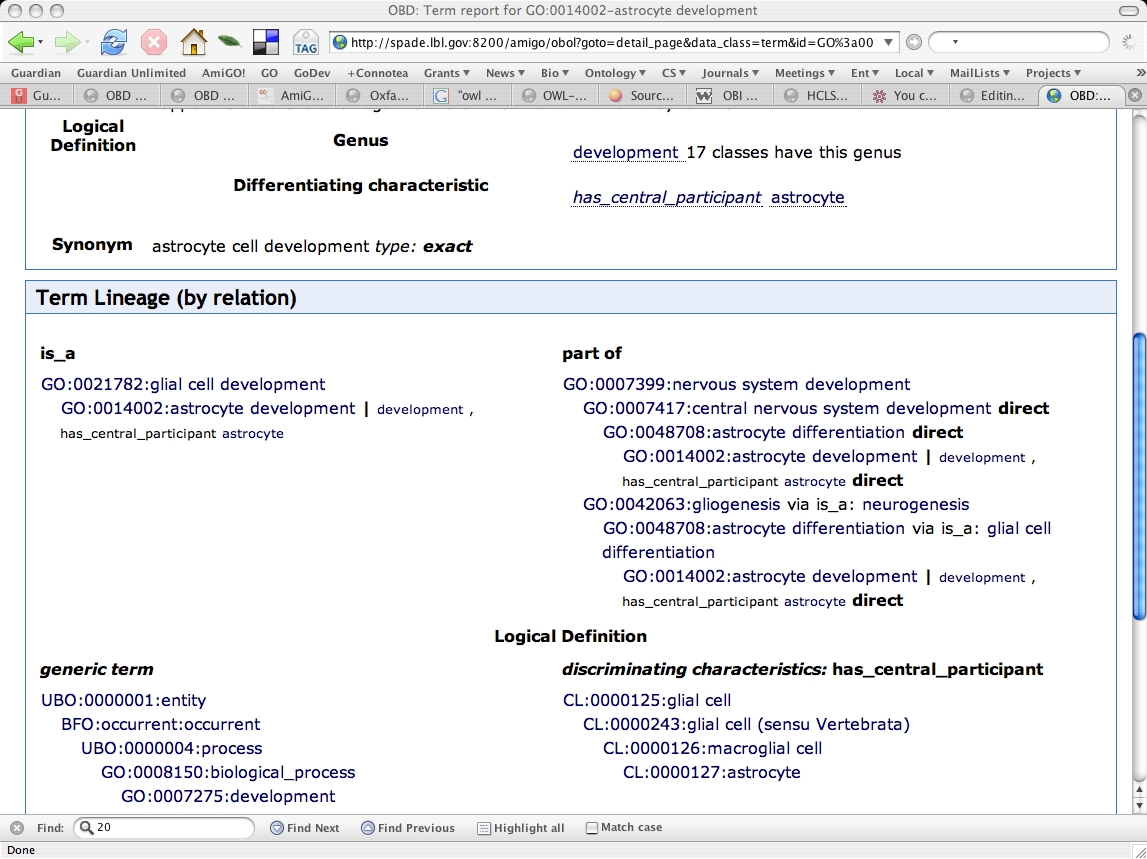
Genus-differentia display
The logical definition is displayed:
Implementation
The browser is implemented entirely in SWI-Prolog using blipkit. The UI borrows heavily from AmiGO, and uses the same CSS code written by Amelia Ireland