Regulation cross-products
For more background, see this presentation:
Many of the items brought up on this page have been resolved.
You should instead look here:
The remainder of this page is out of date
The following has been resolved since this page was created:
- We use 3 relations, regulates, negatively_regulates and positively_regulates
- regulates is transitive_over part_of
Regulation cross-products
The oboedit reasoner can be used to automatically manage regulation terms. This requires the generation of Internal_cross_products. We can do this semi-automatically, because the regulation terms have a consistent simple syntax.
this can serve as a trial run before implementing cross-products with external ontologies
As a first step we only consider Regulation of biological process. For ongoing work see also:
- Regulation of function]
- Regulation of biological quality]
- XP:biological_process_xp_multi_organism_process - for interspecies regulation
Resources
You will need the file gene_ontology_xp.obo, from:
- go/scratch directory
(this file should be updated daily from gene_ontology_edit.obo, provided my cron works)
You can load this into oboedit. Turn the reasoner on; then read on for an explanation
Relations used
Cross-product Definitions
oboedit needs the xp definition (necessary and sufficient conditions) of regulation terms to be made explicit rather than embedded in text. We do this using the new oboformat1.2 intersection_of (cross-product definition) tag.
The idea is that we define a term like negative regulation of synaptogenesis as being:
- A negative regulation process that regulates synpatogenesis
This is an aristotelian (genus-differentia) definition. It can also be seen as the cross-product (intersection) of:
- negative regulation of biological process
- things that regulate synpatogenesis
(This also involves introducing a new relation, regulates)
Here is what this looks like in oboformat:
[Term] id: GO:0051964 name: negative regulation of synaptogenesis namespace: biological_process def: "Any process that stops, prevents or reduces the frequency, rate or extent of synaptogenesis, the formation of a synapse." [] is_a: GO:0051961 ! negative regulation of nervous system development is_a: GO:0051963 ! regulation of synaptogenesis intersection_of: GO:0048519 ! negative regulation of biological process intersection_of: regulates GO:0007416 ! synaptogenesis
Here we have added two lines. These lines can be safely ignored by obof1.2 unaware parsers; they can be stripped out prior to making public if need be. However, they provide oboedit (or any other reasoner-aware tool, like Protege/SWOOP) with the information required to automatically manage the placement of these terms in the DAG.
Editing and browsing regulation cross-products
Oboedit1.1
You can browse and edit the regulation xps in OE1.1
Just load the gene_ontology_xp.obo file from the scratch directory above
In the oboedit cross-product box, this should look like:
Genus: negative regulation of biological process Differentia: part_of synaptogenesis
Oboedit2.0
oboedit2 has more advanced features to make editing cross-products easier
If you are using OboEdit2, you can you the Cross Product Matrix Editor.
here we see a relative sparse area, around mitosis. Not all combinations are realized. Selecting groups of empty cells and clicking the "Make" button realized the terms in the ontology, placing them correctly in the DAG
we can examine a more densely populated area, such as regulation of lymphocyte; clickin on a non-empty cell shows is_a children (pink) and is_a parents (yellow):
Using the reasoner
open the file in the reasoner, you should be able to see both missing is_a links (blue squiggly lines) as well as redundant links (straight red lines).
(See the presentation above for more up-to-date screenshots)
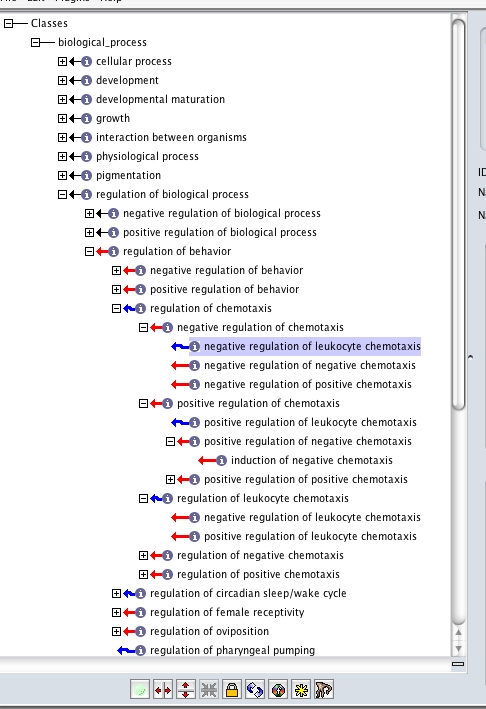
here oboedit is saying that -reg of chemotaxis can be inferred to be an is_a child of -reg of behavior
oboedit can infer this from the logical definition (intersection_of lines). Note that the actual is_a links are not there in the underlying obo file - this is the inferred (implied) graph, not the asserted graph (to see the asserted graph, simply turn off the reasoner; you will see -reg of chemotaxis disappear as a child of -reg of behavior.
Here is another example:
you should be able to see the cross-product definition of the focused term. This term is not asserted to be a child of regulation of cellular process, but this is implied (see the DAG view on the right)
Note the blue lines mean the reasoner knows the link should be there, but it has not been asserted.
In OE2 you can use the "assert implied links" option to fill in missing links:
How was this generated?
How does this all work? I have a simplified version of obol that takes a term
{-/+} regulation of X
And creates a logical definition
genus: {-/+} regulation of biological process
differentia: regulates X
in obo format this is
intersection_of: GO:ID_for_{-/+}_regulation_of_BP
intersection_of: regulates GO:ID_for_X
the oboedit reasoner does the rest
Next steps
We can continue with the process of generating the regulation-xps externally, and periodically running the reasoner to fill in missing links
However, it would be better to start managing the xps directly in the gene_ontology_edit.obo file. New regulation terms would be created using the cross-product interface, and placed in the DAG automatically.
Reasoner details
Given the correct xp definitions, the reasoner can place the regulation terms correctly in the DAG
Skipped links
Not all intermediate links must be filled in the regulation DAG:
Transitivity over part_of
We must have consistent rules about what to do with regulation terms for cases where the regulated stand in part_of relations:
- 2008/01/14 : David/Tanya/Chris : decided regulation is always transitive over part_of. This implies RoMF and RoBP should not be disjoint
Links that cannot be implied
We can also look at regulation is_a links that cannot be implied by the reasoner: